RFLP, RAPD, and AFLP techniques

Introduction:
With the advancement in genetic engineering and diagnostics microbiology, certain molecular or epidemiological techniques have been introduced like RFLP, AFLP and RAPD. Molecular typing techniques play an important role in solving criminal cases and paternity testing and identifying the cause of epidemics or pandemics. Various molecular typing techniques like AFLP, RAPD, and RFLP are employed religiously in diagnostic laboratories and forensic labs. This article describes the separation, isolation, and identification of DNA fragments based on their cleavage site to restriction endonuclease enzyme. RFLP is used in the molecular typing of pathological organisms to identify the source of infection, the cause of infection, and the clonal spread of infection.
RFLP analysis:
Restriction Fragment Length Polymorphism (RFLP) is a technique in which organisms may be differentiated by identifying patterns produced from DNA cleavage. If two organisms have different restriction endonuclease sites on their DNA, the length of the fragments on cleavage produced will differ. The similarity of the patterns generated can be used to differentiate species and even strains from each other.
What are Restriction Endonuclease enzymes?
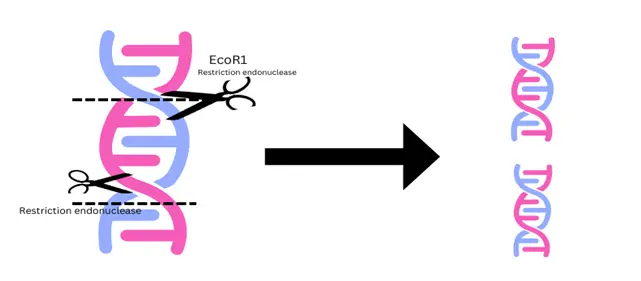
Restriction endonucleases are enzymes that cleave the DNA at specific regions which are usually 10-15 nucleotides in length. The enzyme first recognizes the specific sequence and then cleaves. The shorter the recognition sequence, the greater the number of fragments produced. If molecules differ in the nucleotide sequence, fragments of different sizes may be generated. The fragments can be separated by gel electrophoresis. Restriction enzymes are isolated from various bacteria and are known to play a role in defence mechanisms against the attack of viruses. These enzymes are named by using the first letter of the genus, the first two letters of the species, and the order of discovery.
There are usually two kinds of restriction enzymes based on the length of fragments they produce:
Frequent cutters: The enzymes which produce small and frequent cuts in DNA sequence and produce many small fragments of DNA. For example, EcoR1 from E. Coli strain RY13, and BamH1 from Bacillus amyloliquefaciens.
Rare cutters: The enzyme which cuts rarely in the genome and produces few and large DNA fragments. The sites for rare cutters lie distantly in the genome—for example, Not1 from Nocardia otitis, and Sma1 from Serratia marcescens.
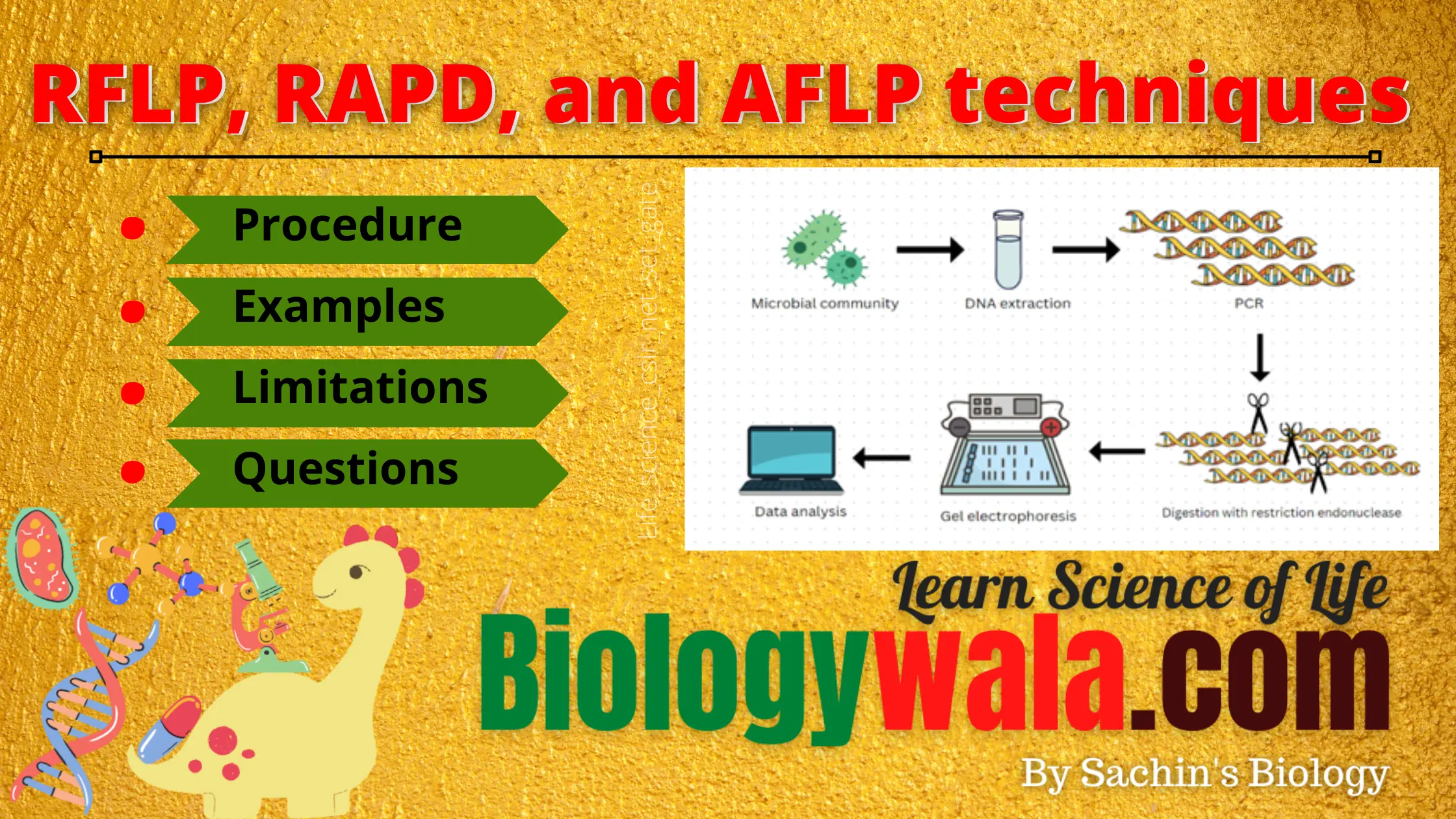
Procedure:
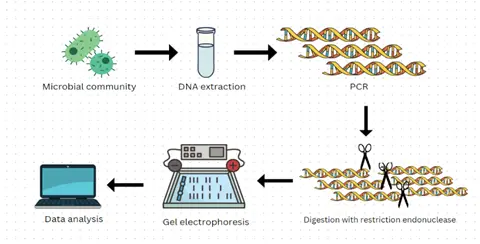
The procedure for RFLP involves the isolation, separation, and identification of DNA fragments. It is a time-consuming and complex process. Sometimes PCR is also engaged when the amount of DNA sample is too small to be analyzed.
Step 1. Extraction: The first step in RFLP is the extraction of the DNA from the sample, (blood, saliva, semen, or some other biological sample or microorganism).
Step 2. Amplification (optional): Amplifying the DNA sample by PCR if the DNA sample is small in amount. PCR is involved in RFLP to amplify the small amount of DNA sample within a few hours and by 20-30 cycles.
Step 3. Production of DNA fragments: The purified DNA is then cut into fragments by the addition of restriction endonuclease enzyme.
Step 4. Gel electrophoresis: The DNA fragments have a negative charge and can be separated by a technique called gel electrophoresis, which separates the pieces of DNA based on their size. The samples of DNA that have been treated with restriction enzymes are placed in separate wells on a slab of electrophoretic gel (agarose gel) across which is placed an electric field. The fragments migrate towards the positive electrode, the smaller fragments moving faster than the larger fragments, thus separating the DNA samples into distinct bands.
Step 5. Detection: The DNA fragments are stained with Ethidium bromide dye while carrying out gel electrophoresis. Ethidium bromide is an intercalating dye that binds to the minor groove of DNA and illuminates when exposed to UV radiation. The orange colour produced by dye helps in detecting the bands of DNA produced of different sizes.
Applications of RFLP:
- RFLPs can be used in paternity cases or criminal cases to determine the source of a DNA sample. (i.e., it has forensic applications).
- RFLPs can be used to determine the disease status of an individual.
- RFLPs can be used to measure recombination rates, leading to a genetic map with the distance between RFLP loci measured in centimorgans.
Example:
1. Identification of cystic fibrosis inheritance from homozygous or heterozygous parents.
- Cystic Fibrosis is a recessive disease, anyone with cystic fibrosis must be homozygous for disease alleles. This information provides us knowledge about the affected and carrier individual in the family.
- For couples expecting a child, it would be simple to test both parents and make a prediction about the eventual disease status of their fetus.
- If both parents are homozygous (wt), then all of their children would also be homozygous.
- If both parents were heterozygous, they could have children with any of the three genotypes, though heterozygous children would be twice as likely as either of the homozygous genotypes.
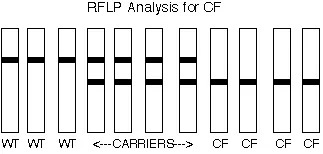
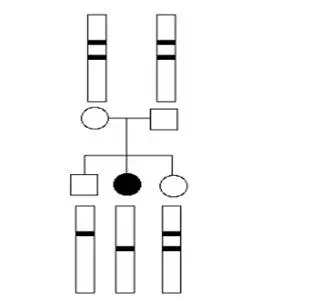
Normal, affected and carrier individual.
2. Determination of Sickle cell anaemia disease.
Sickle-cell anaemia is the result of the change in the 6th amino acid of the beta-globulin chain of haemoglobin where glutamic acid is converted into valine. This results in a change in the shape of RBC from oval to sickle, which leads to multiple problems like oxygen deficiency, and anaemia. The RFLP technique can be used to differentiate between the bands produced by normal RBC and Sickle RBC.
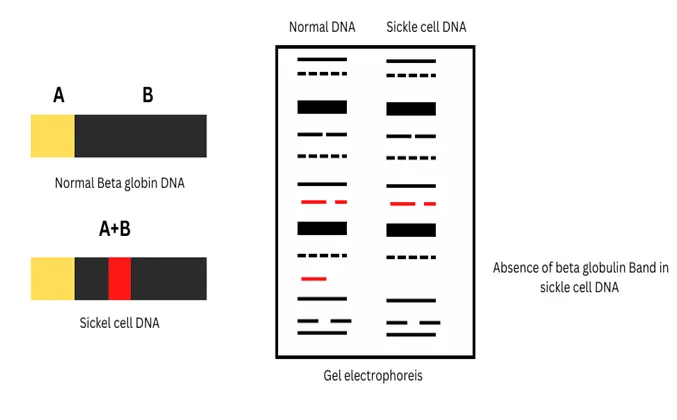
RAPD: Random Amplification of Polymorphic DNA
Random Amplification of Polymorphic DNA is a type of PCR technique that involves the amplification of unknown random DNA fragments using a known single probe. The technique is based on the identification of unknown fragments of target DNA using a known primer. The primer used is short (8-12 nucleotide) arbitrary, and will bind any random fragment of DNA that is complementary to the primer, the bound fragment will be amplified using PCR and then identified, and the triple-stranded DNA is formed as a result of binding to complementary region. This technique is used to determine the genetic diversity among organisms.
No prior knowledge of DNA sequence is required, the primer will bind to the sequence complementary to it but not specified where. The method is beneficial in identifying the target sequence of DNA known to cause disease.
Limitations of RAPD:
- Less resolving power.
- Less reproducibility.
- The mismatch between the primer and DNA sample may alter the results.
- RAPD primers are dominating in nature, so difficult to predict whether the results produced are homozygous or heterozygous.
Procedure:
The RAPD technique works on the principle of binding of short oligonucleotide primer to many different loci in a DNA sample, then the random fragment of DNA is amplified, which is visualized to determine the chromosomal rearrangements or nucleotide variation in the DNA sample.
Step 1. Extraction of DNA sample: The DNA from any source to be analyzed is extracted using various methods.
Step 2. A short, oligonucleotide of known sequence is added to the solution containing the DNA sample.
Step 3. PCR is performed to amplify the bound primer and target sequence of DNA by subjecting it to alternating cycles of renaturation and denaturation. The amplification will take place to only the sequence of DNA with bound oligonucleotide primer with it.
Step 4. The amplified DNA is subjected to gel electrophoresis. The DNA is stained with Ethidium bromide.
Step 5. The electrophoresed DNA will be visualized under a UV illuminator. The distinct bands of amplified DNA will be visible.
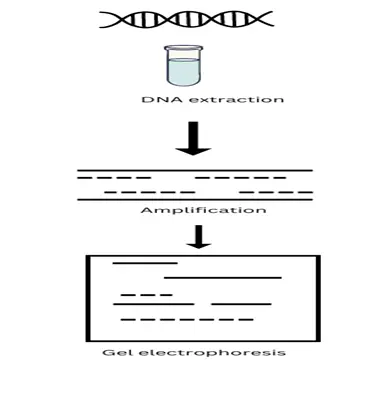
Applications of RAPD:
- Molecular ecology to determine taxonomic identity.
- Assess kinship relationships.
- Analyze mixed genome samples.
- Create specific probes.
AFLP: Amplified Fragment Length Polymorphism
Amplified Fragment Length Polymorphism is another PCR-based technique used in molecular typing and genetic engineering. The technique is based on the binding of complementary adapter molecules to the sticky ends produced after the digestion of DNA with a restriction endonuclease, the bound fragments are then identified by complementary primers and then amplifying the bound fragments.
AFLP not only has higher reproducibility, resolution, and sensitivity than other techniques, but it also can amplify between 50 and 100 fragments at one time.
Adapters? Adapters are short, single-stranded, or double-stranded oligonucleotides used in genetic engineering which can easily bind to sticky ends produced in DNA or RNA and forms the basis of their identification.
Like RAPD, no previous knowledge of DNA templates is required. AFLP requires only a small amount of DNA sample for carrying out analysis. AFLP is a multiplex technique, 100’s fragments can be analyzed in one reaction but it is labour-intensive.
The success of AFLP reaction lies in the use of reagents, optimized reaction conditions and standard and reliable electrophoresis.
Procedure:
The procedure of AFLP involves the production of restriction fragments followed by ligation to the adapter, amplifying the ligated fragment of DNA and then binding with the known primer and selective PCR followed by electrophoresis and detection.
Step 1. Extraction of DNA sample and template preparation.
Step 2. Add the restriction endonuclease enzyme to the DNA solution and incubate.
Step 3. Add the complementary and selected adapter molecule to the reaction mixture and incubate to allow ligation of the adapter to the sticky ends produced due to the digestion of the DNA sample.
Step 4. PCR amplification of the DNA fragments bounds with an adapter i.e., selective PCR using the Taq DNA polymerase.
Step 5. The binding of the amplified DNA fragments with the known primer forms the basis of the identification of the target sequence and then selective PCR to reduce the number of unbound DNA fragments.
Step 6. Separation and visualization of fragments via gel electrophoresis and fragments are stained with Ethidium bromide. The stained DNA fragments are visualized under a UV illuminator.
Applications of AFLP:
- To identify the various polymorphism in the population of an organism.
- To identify the genetic variations of strains and closely related species of organisms.
- In criminal cases and paternity testing.
- Phylogenetic study.
- Physical and genome mapping.
- DNA fingerprinting.
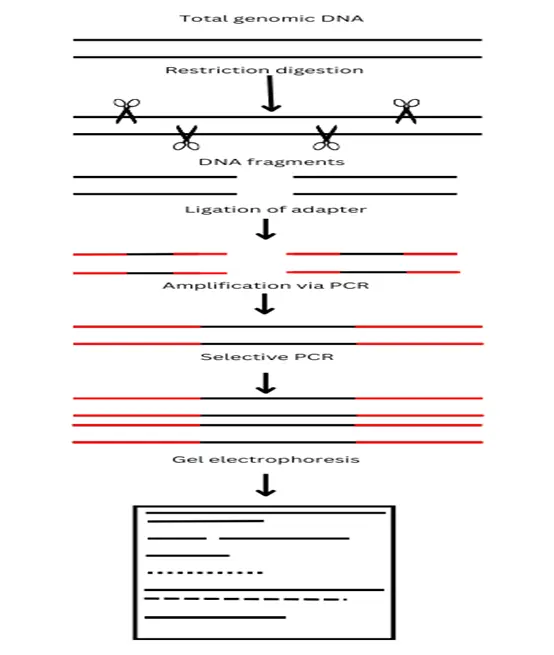
Important questions?
Q.No.1: Difference between RFLP, RAPD, and AFLP?
The major difference is that RFLP is the simplest technique among the three. RFLP requires previous knowledge of target DNA whereas RAPD and AFLP don’t. AFLP employs the use of specially synthesized adapter molecules, and RAPD requires the use of arbitrary oligonucleotide primers of known sequences but RFLP doesn’t employ such complex molecules. RFLP is more commonly used in commercial practice than the other two techniques.
Q.NO.2: How RFLP helps in paternity testing?
RFLP provides the different banding patterns of individual DNA fingerprints and identifying the distance between the cleavage sites and the length of fragments obtained forms the basis of identification. Bands with similar lengths and sizes belong to individuals with similar DNA.
Significance of RFLP and AFLP in genome mapping?
RFLP and AFLP help in locating the gene loci by mapping the distance from restriction endonuclease sites. Which provides knowledge about how far and close the target loci or gene sequence is.
How RAPD helps in epidemiological typing?
RAPD is a molecular typing technique that helps identify the source of infection, and clonal spread by identifying and matching the banding patterns obtained from suspected pathogens and pathogens obtained from infected individuals.
Hope you liked this article about RFLP, RAPD, and AFLP techniques. If you like the article you can let us know on Instagram. We will send you anyhow.
Also like:
[DOWNLOAD] Bruce Albert’s Molecular Biology of The Cell book PDF
[Download] The Biology of Cancer Weinberg PDF Book 2nd edition
[Download] Top 5 Books for Molecular Biology: Basic to Advance
Download Free Reference Books for Genetics
If you want important notes and updates about exams on your mobile then you can join SACHIN’S BIOLOGY on Instagram or Facebook and can directly talk to the founder of Sachin’s Biology and Author of biologywala.com Mr.Sachin Chavan M.Sc. NET JRF (AIR 21) GATE!