Role of chromatin in gene expression and Gene silencing
In the article Role of chromatin in gene expression and Gene silencing, we will be discussing chromatin modifications, Gene silencing, Histone Phosphorylation, Histone acetylation in detail with types of chromatin reorganization by nucleosome remodeling complex.
What is Role of chromatin in gene expression and Gene silencing?
- Chromatin is a genetic material comprising of DNA or RNA and proteins, such as histones. The histones (H1, H2A, H2B, H3, H4) in chromatin helps in the packaging of DNA into structural units called a nucleosome.
- For the packaging of DNA within the nucleus, 146 bp are wrapped around a histone octamer which first recruits two histones H3 and H4 dimers to form a tetramer, and then two histones H2A and H2B dimer.
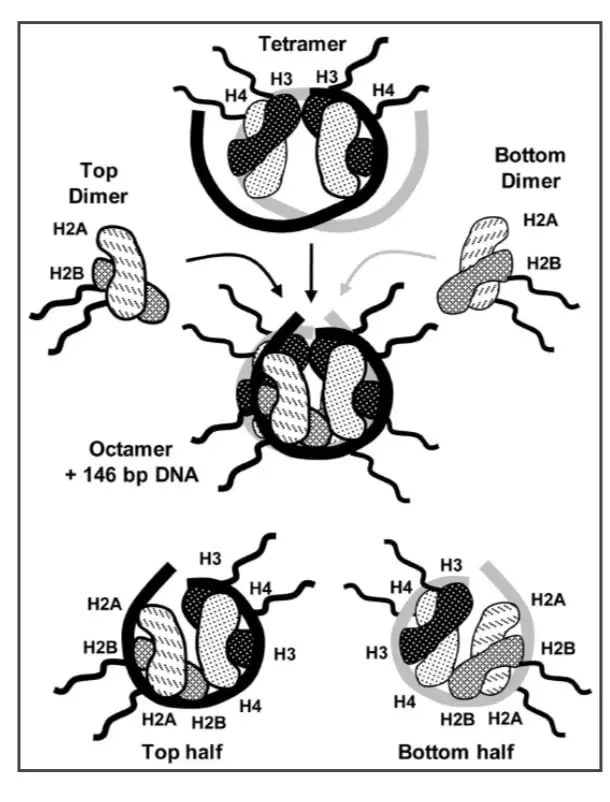
- Chromatin also regulates the expression of the gene in a cell, as compaction of chromatin limits gene accessibility for transcription factors.
- There are two types of chromatin i.e.1)Heterochromatin – chromatin is tightly packed and not actively transcribed. 2)Euchromatin – chromatin is loosely packed and accessible for transcription.
- Thus, for the occurrence of gene expression, changes in chromatin structure take place called chromatin remodeling. These changes are brought by biochemical modifications to DNA and histones including methylation, acetylation, and phosphorylation.
- These modifications depend on the “open” or “closed” state of chromatin which is detected by DNA sensitivity to DNAse I.
Following are the chromatin modifications for the regulation of gene expression:
To understand the role of chromatin in gene expression and Gene silencing, it is necessary to look after 5 main points. These are:
1.DNA methylation:
The structure of DNA can be modified by covalent attachment of methyl(CH3) group called DNA Methylation.
- In eukaryotes, DNA Methylation occurs by an enzyme called DNA methyltransferase (DNMTs), which attaches a methyl group to the 5th cytosine base, forming 5 – methylcytosine.
- DNA Methylation inhibits the transcription of eukaryotic genes when it occurs in the region of the promoter. Near many promoters of genes, CpG island is present.
- CpG site is a location within a DNA sequence in which cytosine and guanine appear consecutively. (5’C-P-G-P-C-P-G3′)
- CpG islands are commonly 1000 to 2000 Bp in length and contain a high number of CpG sites. These CpG islands are called genetic hotspots because these are the regions for active methylation.
- DNA methylation plays important role in the silencing of tissue-specific genes to prevent them from being expressed in the wrong tissue.
- When CpG island in a promoter region is methylated certain proteins bind to it which blocks the binding of transcription factors and inhibits the transcription.
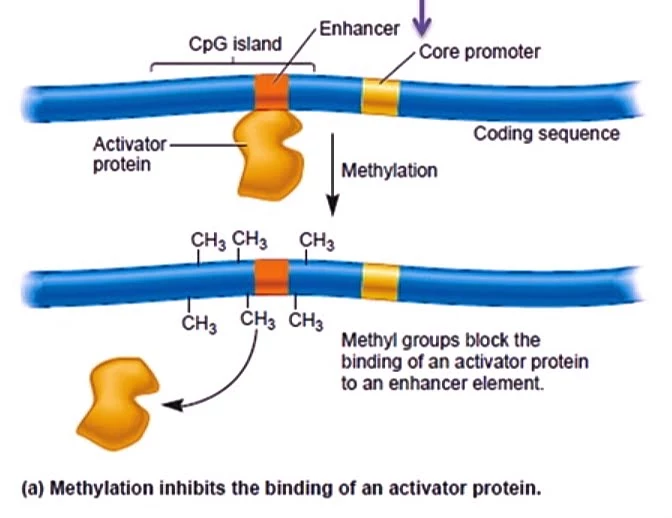
- There are some other sequences which present in DNA that can also act as gene silencers called Gene insulators.
- An insulator is a segment of DNA that acts as a boundary between two genes. It insulates or protects a gene from the regulatory effects of neighboring genes. For example, as given in the diagram below if gene ‘A’ is transcribed, the enhancer will be produced, and this enhancer will bind to the insulator region. As it is bound to the insulator region both the expression of gene ‘A’ and gene ‘B’ will occur independently.
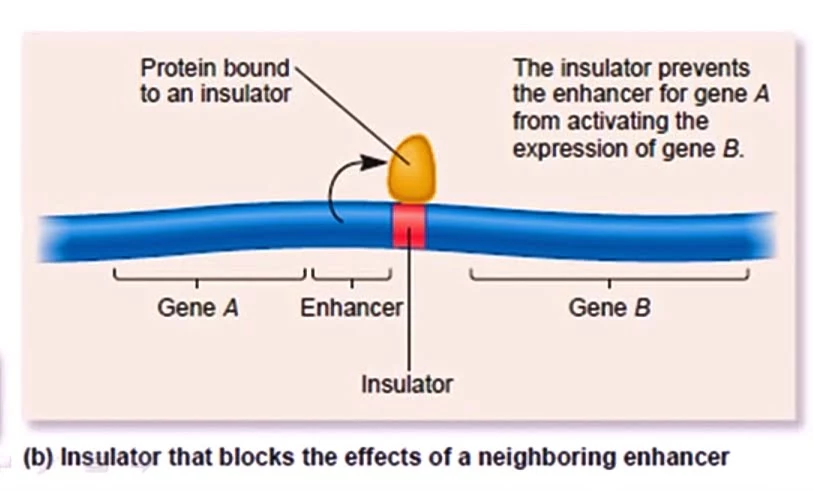
- Gene insulator plays a major role in genomic imprinting. Genomic imprinting is a phenomenon in which the expression of certain genes is determined by whether the gene is maternal or paternal i.e. inherited from the female or male parent.
- For example, the gene Igf2 is an imprinted gene and is expressed only from the paternal chromosome. Similarly, gene H19 is an imprinted gene expressed only from the maternal chromosome.
- Gene Igf2 and H19 both present on maternal as well as paternal chromosomes. But these genes are separated by an insulator sequence. The protein CCCTC – binding factor (CTCF) bind to this insulator sequence and makes a wall that inhibits the expression of one gene which is not necessary and allows the expression of another gene which is a specific imprinted gene for that maternal or paternal chromosome.
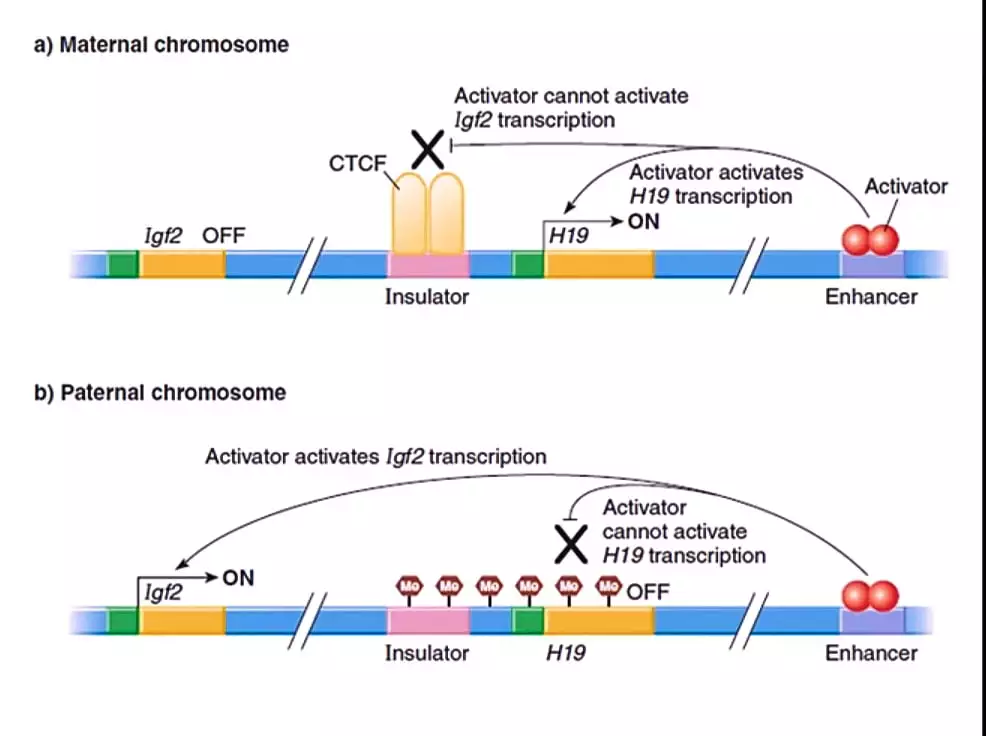
- In a nutshell, due to the presence of an insulator sequence and a binding protein (CTCF), the paternal chromosome will express the gene Igf2 but not the gene H19. Similarly, in the maternal chromosome, the gene H19 will express but not the gene Igf2. This controls the gender or sex-based expression of genes.
2. Histone methylation :
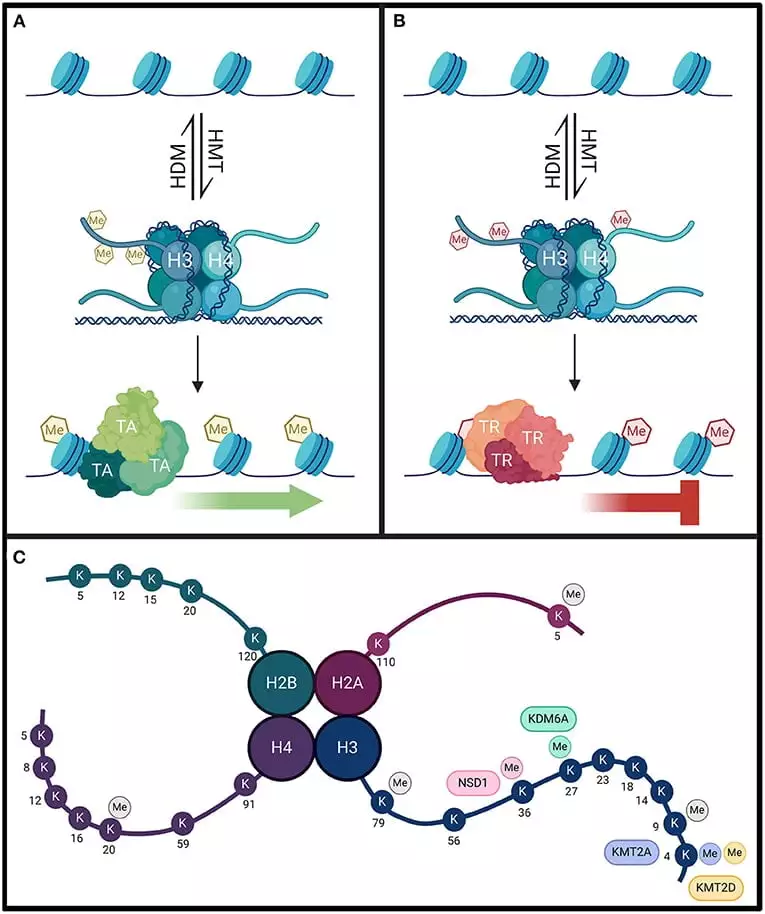
- In this process methyl group (CH3) is added to lysine or arginine via the enzyme Histone methyltransferase (HMT) with the help of a donor s- Adenesyl methionine. This is a covalent modification. Histones have shown that several lysines residues including lysine 4,9,27,36bof of H3 and lysine 20 of H4 are preferred sites of methylation. Histone methylation exists in mono, di, tri methylation forms.
- When the methyl group is attached to the 4th and 36th positions of lysine transcription gets activated and when attached to the 9th and 27th positions it inhibits the transcription.
- As it is a reversible process so demethylation occurs by histone deacetylase (HDM).
- Histone methylation determines the expressed or repressed state of the surrounding genes by altering the chromatin structure.
3. Histone acetylation :
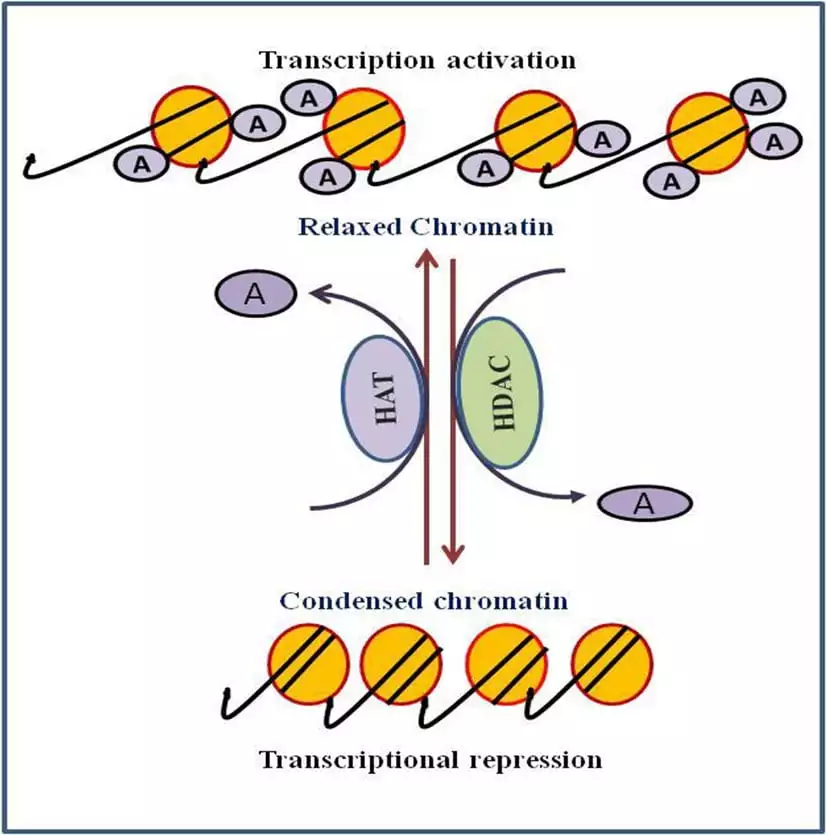
- Histone acetylation and deacetylation play an important role in regulating gene expression making genes more or less available for transcription. In histone acetylation, the acetyl group is added usually to lysine residues at the N- terminus by histone methyltransferase (HAT).
- As histones are positively charged, when HAT protein attaches acetyl groups to these histones the positive charge gets neutralized. As a result DNA(having a negative charge) is less attracted to histones and bonds between DNA and histones get weakened.
- While in histone deacetylation acetyl group is removed from lysine by histone deacetylase (HDACs) which allow histones to wrap DNA more tightly and block the transcription.
4. Histone Phosphorylation :
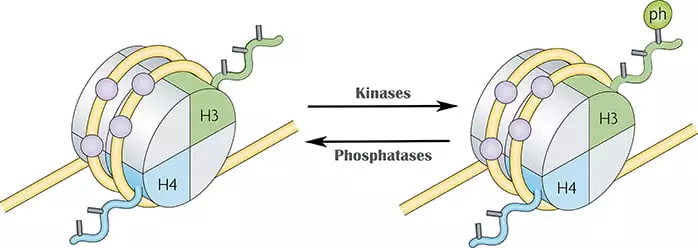
- Phosphorylation occurs on serine, tyrosine, and threonine residues on the N- terminal histone tails. It attaches the phosphate group by enzyme kinase which confers a negative charge to histone as result chromatin gets open.
- Histone phosphorylation occurs in DNA repair, cell division, apoptosis, and transcription activation. Phosphorylation of serine(Ser) 10 and 28 on H3 and 32 on H2B is associated with the activation of gene transcription.
- Tyrosine phosphorylation is associated with transcription initiation. The linker histone H1 stabilizes nucleosome is also phosphorylated called as H1 phosphorylation which leads to chromatin relaxation to allow transcription. During meiosis and mitosis histone phosphorylation plays a major role.
5. Histone ubiquitylation :
- In this modification process, the ubiquitin-protein is covalently linked to the lysine residue or target proteins.
- The bond between a lysine residue and ubiquitin-protein is an isopeptide. The purpose of ubiquitination is to mark the proteins for destruction or degradation.
- The ubiquitination occurs on that protein, which is misfolded or toxic. Ubiquitination happens to be part of some signaling pathways to switch on and switch off the pathway by the destruction of proteins.
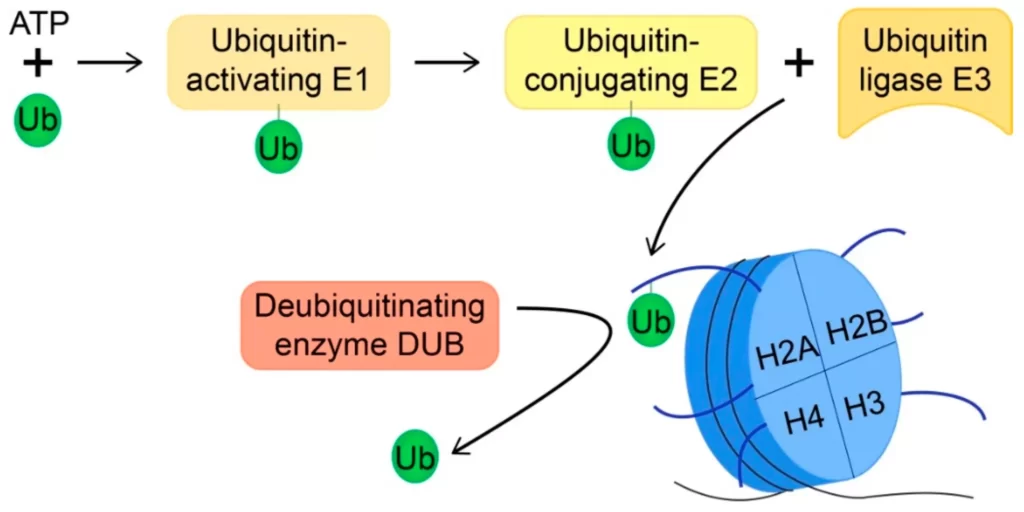
- Enzymes involved in the ubiquitination process: 1) Ubiquitin-activating enzymes (E1) – This enzyme links itself with the carboxyl end of the ubiquitin enzyme through the Thioster bond. As this step is energy-dependent it needs ATP. 2) Ubiquitin-conjugating enzyme (E2) – It transports the activated ubiquitin from E1 to cysteine residue of the E2 enzyme. 3)Ubiquitin-ligase enzyme (E3) – Its function is to transfer the ubiquitin from E2 to the target protein.
Types of chromatin reorganization by nucleosome remodeling complex :
Nucleosome sliding –
- Nucleosome sliding is catalyzed by ATP-dependent nucleosome remodeling factors. These factors modulate the structure of chromatin in more subtle ways. the remodeling factors use the freed by ATP hydrolysis to disrupt the DNA -histone interactions. Their action increases the accessibility of nucleosomal DNA and also facilitates the relocation of the histone octamer.
- The histone octamer relocates along with DNA without dissociating from it. Nucleosome sliding occurs through two mechanisms i.e. 1) Twist diffusion and 2) Loop /bulge propagation.
- In twist diffusion, a single base pair can be transferred between DNA and histone core. As a result, the segment of nucleosomal DNA twists or untwist to accommodate the loss or gain of that base pair. If a twist from one base pair were propagated all around the nucleosome from one DNA segment to the next it is called a twist diffusion.
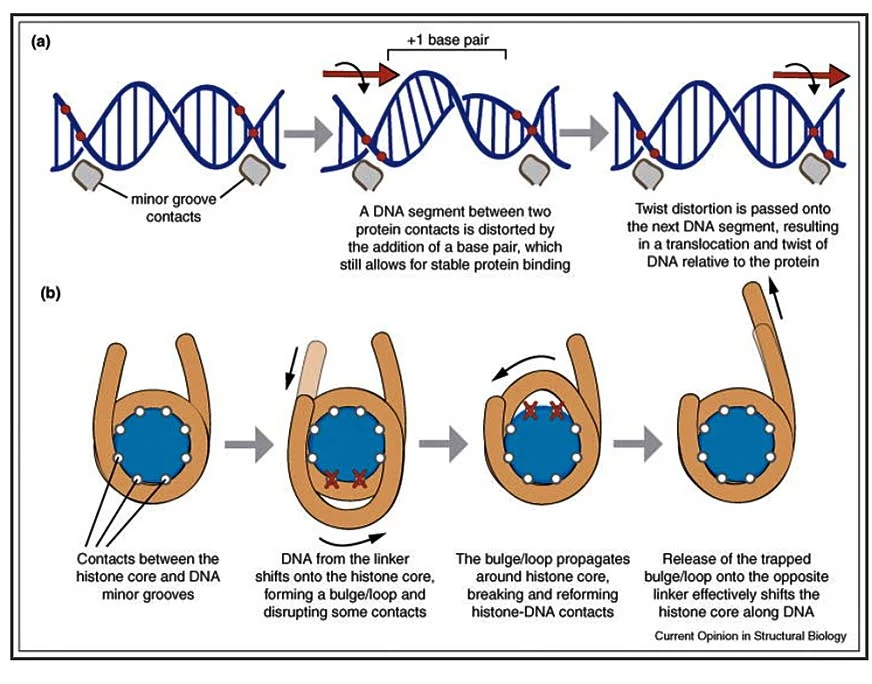
- Like the twist diffusion, loop/bulge propagation also suggests that when DNA from one linker shifts on the nucleosome creates a loop or bulge of DNA which diffuses around the histone core to emerge on another side.
- Due to the process of nucleosome sliding, there is a change in internucleosomal space which allows the DNA polymerase to bind and start transcription which eventually leads to the expression of genes.
2. Histone eviction:
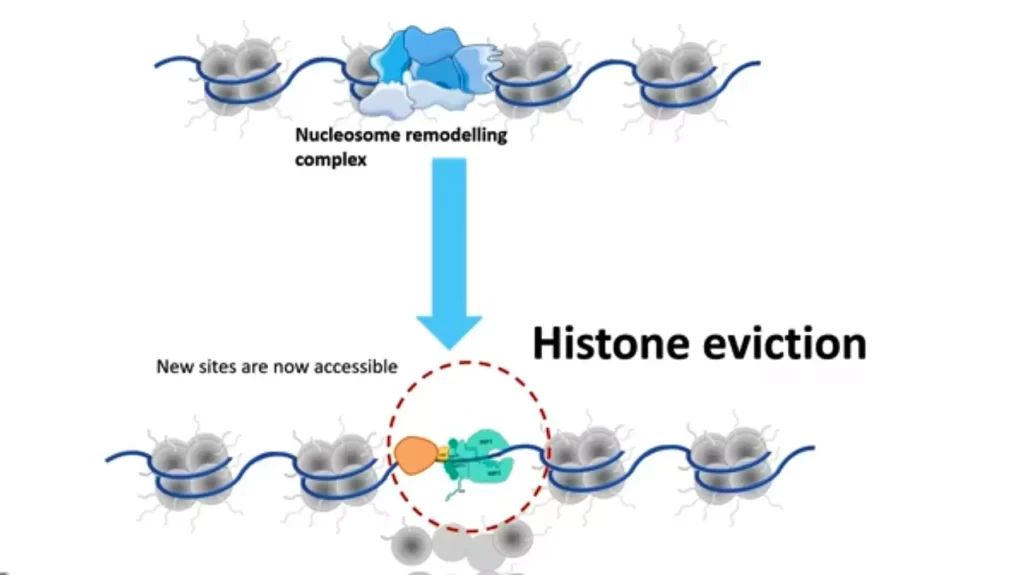
- Histone octamers which are wrapped around DNA can be evicted by the nucleosome remodeling complex which allows the release of the nucleosome core and frees up DNA for interacting with other DNA binding proteins and repair machinery.
- After histone eviction, the new sites which are accessible can bind to transcription factors and allow gene expression.
3. Histone replacement:
- Histone is replaced by different variants of histones by using a nucleosome remodeling complex. Let’s consider, there is a double-stranded DNA break, to repair this break the repair machinery needs to be recruited while it is in a compact form. So the nucleosome remodeling complex loosens the structure and makes it more accessible to recruit the repair machinery. The common way how nucleosome remodeling complex does this is by the replacement of histones.
- There are various families in the nucleosome remodeling complex such as SWI/SNF (common in Yeast, Human & Drosophila), ISWI, IN080, M12/Chd, etc. Each of these complexes has an ATPase domain and all of them need ATP hydrolysis for their functioning.
- ISWI remodelers organize nucleosomes into a proper bundle form and create equal spacing between nucleosomes. It also plays a central role in chromatin assembly after DNA replication and maintenance of higher-order chromatin structure. SWI/SNF remodelers disorder nucleosomes.
Gene silencing: RNA interference pathway
- The prevention of the expression of the gene is termed gene silencing. Why do we require to silence or prevent the expression of the gene? In the complete set of genes, some of the genes can have mutations. Due to these mutations, these genes might not function in a normal way. So to prevent the expression of these genes cells need to silence or degrade that gene.
- The silencing of genes is carried out by the RNA interference mechanism(RNAi). RNA interference is used to degrade target DNA by utilizing a different mechanism like siRNA(short interfering RNA) and miRNA (microRNA).
siRNA mechanism :
- Small/short interfering RNA or silencing RNA is a double-stranded, non-coding RNA molecule. These RNA molecules are 19-23bp in length and function within the RNA interference pathway by degrading mRNA and preventing the synthesis of target proteins from mRNA.
- So where do these mRNAs come from? The siRNA is exogenous in origin means they come outside of the cell. Cell uptake this double-stranded siRNA from its surroundings and it will process that ds RNA into ss RNA and form a complex to degrade the protein.
- The mechanism starts with either ds RNA or hairpin structured RNA. The hairpin loop is cleaved by the enzyme Dicer and form siRNA duplex.
- As siRNA duplex is formed, the protein called Argonaute(Ago) load itself with some other accessory proteins and form RISC(RNA Induced Silencing Complex). Among the two strands of siRNA, only one strand is used which is called the Antisense strand. This antisense strand has a sequence similarity to bind with the target mRNA.
- Along with the antisense strand, the RISC complex loads itself to the target mRNA called siRNA-mRNA complex.
- The siRNA -mRNA cleaves the target mRNA and as a result, mRNA gets spliced or degraded. In the first step where the river cleaves the RNA, ATP is utilized as an energy source as well as during the separation of strands for the RISC recruitment there is also a need for ATP as an energy source. Both these stages are ATP-dependent.

What are RISC complex and Argonaute protein?
- Argonaute protein, DGCR8(accessory protein), or sometimes the dicer together forms a RISC complex.
- RISC is an RNA multi-protein complex that degrades the mRNA in response to siRNA.
- The active components of RISC are endonucleases called Argonaute proteins which cleave the target mRNA strand. Argonaute protein plays a vital role in the RISC complex. It has two domains called the PAZ domain (Piwi/Argonaute&Zwille) and the PIWI domain (P-element Induced Wimpy Testis).
- PAZ domain binds to nucleic acid and recognizes 3′ end of ss RNA and PIWI domain districts the phosphodiesterase bond and cleaves the target mRNA.
2. miRNA mechanism:
- A micro RNA is a small non-coding RNA molecule that functions in RNA silencing and transcriptional regulation of gene expression. The double-stranded RNA can be converted into siRNA or miRNA. Whenever siRNA or miRNA is formed it will ultimately degrade the mRNA and inhibit protein synthesis.
- miRNA is mostly endogenous in origin because the miRNA will be produced by the cell itself in the nucleus and not taken from the surrounding. Most of the miRNA is produced from the non-coding region is called introns and less from exons and the length is about 19-24bp.
- There are two stages of miRNA mediated pathway. One stage happens in the nucleus and the other is in the cytoplasm.
- A long stretch of siRNA is produced in the nucleus which has self-complementary regions so that they can bind with themselves and form a loop-like structure called pri-miRNA. This pri-miRNA gets converted into pre-miRNA by the enzyme Drosha. Enzyme drosha cleaves pri-miRNA and converts it into pre-miRNA.
- This pre-miRNA is then transported to the cytoplasm via a special export receptor present in the nuclear envelope known as Exportin 5.
- In the cytoplasm, the Dicer enzyme cleaves the pre-miRNA.(cleaves the loop made by self-complementary regions and form miR/miR duplex)
- This miR/miR start duplex gets converted into mature mRNA by the RISC complex. The RISC complex removes one of the strands and takes one single strand along with argonaute protein and then binds to the target mRNA and degrade it.
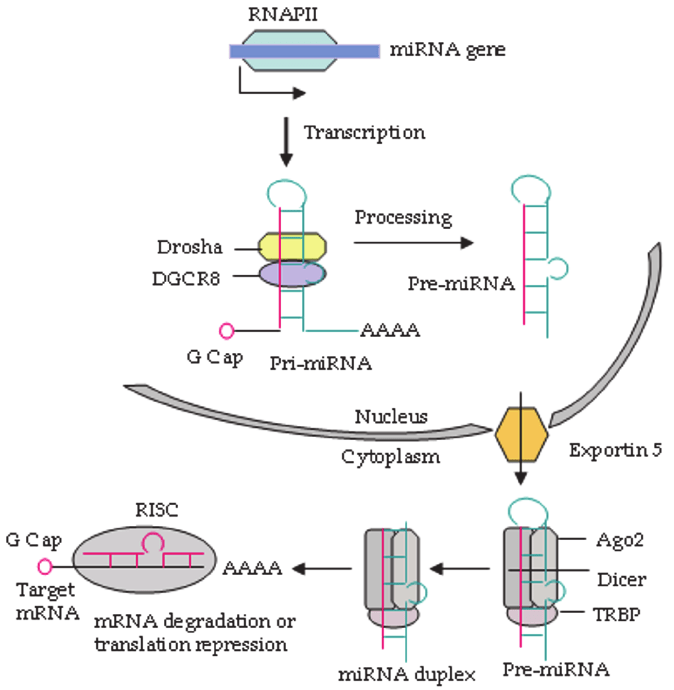
- The major difference between siRNA and mRNA mechanisms is that mRNA needs one extra enzyme called Drosha.
- siRNA → Exogenous origin →Dicer
- miRNA→Endogenous origin→Dicer , Drosha
Frequently asked questions about the Role of chromatin in gene expression and Gene silencing:
How does chromatin structure affect gene expression?
In eukaryotes, regulation of gene expression can be controlled by tight or loose packaging of genes in chromatin(DNA & histones). The structure of chromatin can be altered by various types of modifications in DNA and Histones such as methylation, acetylation, phosphorylation, etc. Various chromatin modellers also help in altering the structure of chromosomes. The compaction of DNA in the form of chromatin limits gene accessibility for transcription factors. Different modifications like nucleosome sliding, histone eviction, histone replacement modify the chromatin structure and make it available for gene expression.
What is gene silencing and how it is carried out?
Gene silencing is the prevention of the expression of the gene. When a mutant gene is present in a set of genes, to prevent the malfunction of the mutant gene, cells need to silence or degrade that gene. Gene silencing works either by suppressing transcription or by the RNA degradation process (Transcriptional gene silencing[TGS] and Post-transcriptional gene silencing or RNA interference[RNAi]). TGS mechanism is different from the RNAi mechanism. The consequences of TGS are long-term that can pass on to the daughter cells during cell division, which is not the case in RNAi.
What is the RNAi mechanism?
RNA interference is the mechanism used to degrade target RNA by utilizing different mechanisms like siRNA(Short interfering RNA) and miRNA (microRNA). RNAi mechanism regulates gene expression by inhibition of translation of mRNA or by direct degradation of mRNA. This mechanism is necessary for defence against the infection by viruses, protection from transposons. By using non-coding RNA molecules like siRNA & miRNA, RNA induced silencing complex (RISC) is formed that cleaves or degrade the mRNA by attaching to the target mRNA with the help of enzymes Dicer, Drosha. Both siRNA and miRNA are distinct from where they originate. siRNA is exogenous in origin although miRNA is endogenous. siRNA is 100% complementary to the target but miRNA originates with ssRNA forming a hairpin structure which is why not 100% complementary to the target. The enzyme Dicer works in siRNA while miRNA needs one extra enzyme called Drosha along with Dicer.
Hope you liked this article about the role of chromatin in gene expression and Gene silencing. If you think there is something in this article or need any kind of improvement you can let us know in the comment box below. After reading the Role of chromatin in gene expression and Gene silencing you will also like other articles on molecular biology.
You also like:
Regulation of gene in Prokaryotes
[Download] Free Molecular Biology of Cancer PDF Book 4th Edi.
If you want important notes and updates about exams on your mobile then you can join SACHIN’SBIOLOGY on Instagram or Facebook and can directly talk to the founder of Sachin’s Biology and Author of biologywala.com Mr Sachin Chavan M.Sc. NET JRF (AIR 21) GATE!
Take the 1st step of your CSIR NET JRF Life Science preparation journey by attempting one of these 5 quizzes:
![[DOWNLOAD] GATE Life Science Question Paper PDF FREE 15 [DOWNLOAD] GATE Life Science Question Paper PDF](https://biologywala.com/wp-content/uploads/2023/04/we-will-provide-a-comprehensive-guide-on-downloading-GATE-Life-Science-question-paper-PDFs-and-how-to-use-them-for-effective-preparation-520x245.webp)
